or the Magic Mushrooms Keychain
“Magic mushrooms” are fungi that contain Psylocibin.
When we ingest Psylocibin, it gets degraded by the acid juices of our stomachs and loses its phosphate group (P), giving rise to a compound called Psilocin.
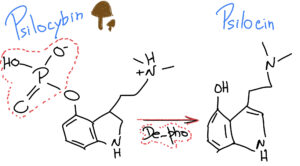
What is interesting is that Psilocin (organic name: 4-hydroxy-N,N-dimetiltryptamine) is very similar to Serotonin – a very important neurotransmitter involved in our mood, learning and a plethora of other fundamental physiological processes.
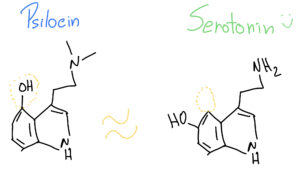
As such, when Psilocin reaches our prefrontal cortex, it can easily bind to our serotonin receptors, because they look so similar – it’s like two old keys that look almost alike and can open the same door at our grandmother’s house.
But Psilocin and Serotonin are indeed different.
As such, Psilocin is able to do certain things in our brains when it binds to the similar Serotonin “door lock”, causing hallucinations and emotional changes that seem to alter the perception of space and time. Psilocin is like that funny cousin that can create chaos when it comes to visit during summer vacations… And, not all trips seem to be good trips, because as all things in life, a lot depends on the surrounding environment and the dosage. So, if a person comes through that door in a poor state of mind, it will just go down the “dark-hole” even further – so they say…
In fact a couple of years back, psychopharmacologists Robin Carhart-Harris and David Nutt from the Imperial College London did a fMRI study (functional magnetic resonance imaging) to evaluate the effects of Psilocybin in the brain2, and decided to give it IV (intravenously) to quicken the trip-effect because they were scared the “voyage” inside of the tight-noisy fMRI machine could be scary for the 30 individuals high on Psilocybin3. The results were quite interesting and showed that the effects of this psychedelic drug could be caused by a decreased activity and connectivity in the brain’s key connector hubs, like enabling a state of unconstrained cognition3. It seems Psilocybin reduced the blood flow and neural activity in the posterior cingulate cortex and medial prefrontal cortex – almost like making a “software reset” of the brain.
As such, Psilocin has been considered a serotonergic psychedelic compound; and it has been banned since the 70’s because people at the time thought it had no therapeutic value.
But Psilocin seems to have an effect in the treatment of Major Depressive Disorder (MDD), a leading cause of disability worldwide. Robin von Rotz and team at the Neurophenomenology of Conscious Lab from the University of Zürich, Switzerland, have just released the results of a randomized double-blind clinical trial 1. This clinical study showed that a single, moderate dose of Psilocybin (0.215 mg/Kg) significantly reduced depressive symptoms compared to a placebo, the “sugar-pill” that they give to the control group.
Even though the results were only evaluated for a period of two weeks after ingestion, the depression severity scores significantly improved in the treated patients in comparison with controls. So, this is one of the first clinical studies to actually demonstrate improvements directly attributed to Psilocybin/Psilocin itself. If we think that the state of deep depression is actually a neural circuitry disfunction, then Psilocin with its similar key structure to Serotonin might be able to open and clean the faulty neuronal-wires that contribute to the brain disfunction seen in MDD.
Several larger clinical studies are currently under way that could eventually pave the way to full regulatory approval, and the removal of Psilocybin from the banned WHO list of pure psychedelic drugs. The U.S. Food and Drug Administration already gave psilocybin the “breakthrough therapy” designation for MDD and Treatment-Resistant Disorder; and, in Australia some psychiatrist can have permission to use it under certain conditions for Post-Traumatic Stress Disorder. The European Medicines Agency (EMA) is following suit, and its Chief Medical Officer has just released a statement that it is actively engaged with developers of psychedelic therapies and academic researchers to help them identify what it takes to move forward and fully bring psychedelics as medical therapies to our pharmacies (and not street dealers)4.
We will be on the lookout for those results…

References:
1 von Rotz, R. et al. Single-dose psilocybin-assisted therapy in major depressive disorder: a placebo-controlled, double-blind, randomised clinical trial. eClinicalMedicine 56 (2023). https://doi.org:10.1016/j.eclinm.2022.101809
2 Carhart-Harris, R. L. et al. The administration of psilocybin to healthy, hallucinogen-experienced volunteers in a mock-functional magnetic resonance imaging environment: a preliminary investigation of tolerability. J Psychopharmacol 25, 1562-1567 (2011). https://doi.org:10.1177/0269881110367445
3 Miller, G. Mapping the psychadelic brain. Science Brain & Behaviour (2012). https://doi.org:10.1126/article.27824
4 https://www.linkedin.com/pulse/second-chance-psychedelics-european-medicines-agency


