The n-of-1 patient is that one person with a unique genetic mutation that causes an ultra-rare disease, designated as a disease with less than 30 patients in the whole world. The advent of affordable genomic sequencing has identified millions of n-of-1 patients, which is becoming a large and growing population with desperate needs.
Though a great progress is made in identifying n-of-1 patients, and ruling the genetic causes of their diseases; usually, drugs that work in patients with the common mutations, often do not work for these unique patients.
n-Lorem is a foundation created by Dr. Stanley Crooke, the founder, chairman and chief executive officer of Ionis Pharmaceuticals, a global leader in RNA-targeted therapy. This foundation was created to provide individualized treatments for patients with ultra-rare diseases using the technology developed at Ionis Pharmaceuticals.
The mission of n-Lorem is to use the versatility and specificity of antisense technology to kindly offer experimental antisense oligonucleotides (ASO) medicines to treat the n-of-1 patient.
Antisense oligonucleotides (ASOs) are short, synthetic, single-stranded DNA-mimics that can alter the RNA, and consequently protein expression and function. Because they can manipulate the intermediate step between gene and protein translation – at the pre-mRNA level – ASOs can reduce, or restore, or modify protein expression by different mechanisms.
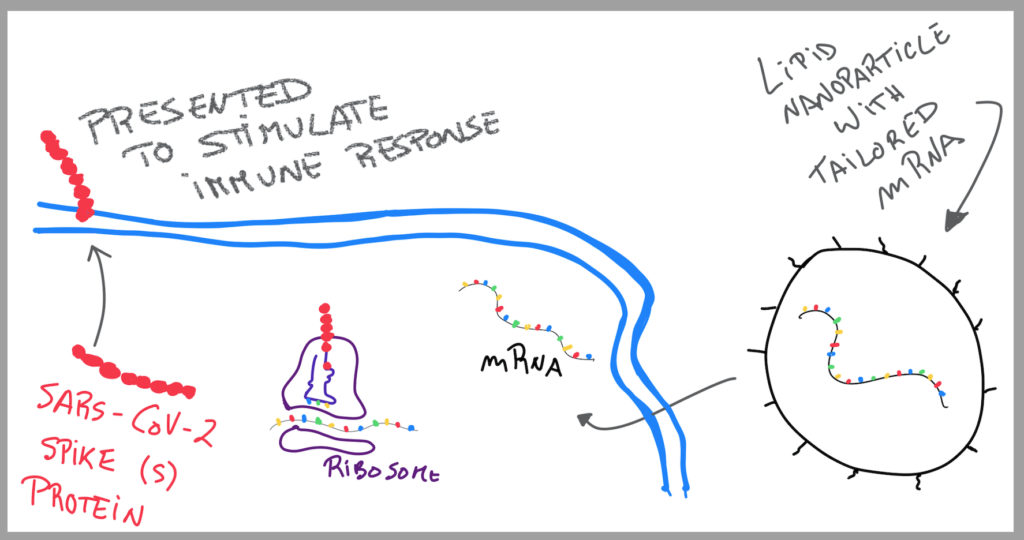
Very simply: DNA gets converted into mRNA. ASOs function in between the two. As such, if the DNA gene has a mutation and codes a dysfunctional mRNA, and consequentially a dysfunctional protein that causes a disease, an ASO can change that. The ASO has the correct complementary code to produce the precise mRNA, and automatically, the precise protein needed to perform the right function. By intervening in the step before protein gets translated, it makes sure that nothing goes wrong and that the mutated gene doesn’t go any further, preventing disease from happening.
Even though it only started one year ago, n-Lorem together with Ionis Pharmaceuticals have helped design and provide experimental ASOs for two unique patients: one with Batten’s disease and ataxia-telangiectasia (AT); the other one, with a FUS mutation in Amyotrophic Lateral Sclerosis (ALS). These have provided a tremendous opportunity to certify the n-Lorem concept and drive motivation forward.
Anyone can apply to n-Lorem for a potential treatment. The proposals are approved and prioritized based on certain criteria, such as the severity of the disease, feasibility of developing an ASO treatment for the genetic cause of the disease, degree of potential benefit vs. potential risks, practicality of treatment, availability of physician and institution to treat patient, and other intricacies of the condition.
The unique patient needs to work with a physician that makes the connection to n-Lorem, who will then make an informed decision about whether a patient is appropriate to receive an experimental ASO treatment through a Commission.
It’s outstanding that the FDA reaction to n-Lorem has been very supportive; and, in fact, initial guidance has been put in place in January 2021 to reach more patients in need.
Once regulatory permission has been given, an investigator-led clinical trial is initiated and the patient can receive their custom experimental ASO treatment at no personal cost and will all clinical support.
This cost-free individualized therapy is possible for Ionis Pharmaceuticals because of the inherent efficiency and versatility of the ASO technology. The knowledge of modern ASOs mechanisms and specific possible effectiveness in selected organs, with different possible routes of application, together with integrated safety databases, allows a dive for the treatment of unique patients.
It’s honourable to use science to help the n-of-1 patient and their families.
I hope for more Dr. Stanley Crooke’s…